Difference between revisions of "PGMS-pathway control state"
From Bioblast
m (Cardoso Luiza moved page PGMS pathway control state to PGMS-pathway control state: Edited to have same nomenclature used in other pages) |
|||
| (7 intermediate revisions by 5 users not shown) | |||
| Line 3: | Line 3: | ||
|description=[[File:PGMS.png|left|200px|PGMS]] '''PGMS''': [[Pyruvate]] & [[Glutamate]] & [[Malate]] & [[Succinate]]. | |description=[[File:PGMS.png|left|200px|PGMS]] '''PGMS''': [[Pyruvate]] & [[Glutamate]] & [[Malate]] & [[Succinate]]. | ||
'''MitoPathway control state:''' NS | '''MitoPathway control state:''' [[NS|NS-pathway control state]] | ||
2-oxoglutarate is produced through the citric acid cycle from citrate by isocitrate dehydrogenase, from oxaloacetate and glutamate by the transaminase, and from glutamate by the glutamate dehydrogenase. If the 2-oxoglutarate carrier does not outcompete these sources of 2-oxoglutarate, then the TCA cycle operates in full circle with external pyruvate&malate&glutamate&succinate | 2-oxoglutarate is produced through the citric acid cycle from citrate by isocitrate dehydrogenase, from oxaloacetate and glutamate by the transaminase, and from glutamate by the glutamate dehydrogenase. If the 2-oxoglutarate carrier does not outcompete these sources of 2-oxoglutarate, then the TCA cycle operates in full circle with external pyruvate&malate&glutamate&succinate | ||
|info=[[Gnaiger | |info=[[Gnaiger 2020 BEC MitoPathways]] | ||
}} | }} | ||
::: ''More details'' | |||
| | ::::» [[NS-pathway control state]] | ||
::::» [[Additive effect of convergent electron flow]] | |||
::::» [[Respiratory_complexes#Respiratory_complexes_-_more_than_five |Respiratory complexes - more than five]] | |||
::::» [[Convergent electron flow]] | |||
== PGMS | == PGMS<sub>''L''</sub> == | ||
:::: PGMS pathway in the LEAK state can be evaluated in the following SUIT protocols: | |||
== PGMS( | == PGMS<sub>''P''</sub> == | ||
:::: PGMS pathway in the OXPHOS state can be evaluated in the following SUIT protocols: | |||
:::*[[SUIT-008]] | |||
::::* DL-Protocol for permeabilized fibers (pfi): [[SUIT-008 O2 pfi D014]] | |||
::::* DL-Protocol for permeabilized cells (pce): [[SUIT-008 O2 ce-pce D025]] | |||
:::*[[SUIT-014]] | |||
== PGMS<sub>''E''</sub> == | |||
:::: PGMS pathway in the ET state can be evaluated in the following SUIT protocols: | |||
:::*[[SUIT-001]] | |||
::::* DL-Protocol for isolated mitochondria and tissue homogenate (mt): [[SUIT-001 O2 mt D001]] | |||
::::* DL-Protocol for permeabilized fibers (pfi): [[SUIT-001 O2 pfi D002]] | |||
::::* DL-Protocol for permeabilized cells (pce): [[SUIT-001 O2 ce-pce D003]] | |||
::::* DL-Protocol for permeabilized PBMC and PLT: [[SUIT-001 O2 ce-pce D004]] | |||
:::*[[SUIT-008]] | |||
::::* DL-Protocol for permeabilized fibers (pfi): [[SUIT-008 O2 pfi D014]] | |||
::::* DL-Protocol for permeabilized cells (pce): [[SUIT-008 O2 ce-pce D025]] | |||
:::*[[SUIT-014]] | |||
== Discussion == | == Discussion == | ||
:::: Recent studies showed that S- and NS-linked OXPHOS capacity is inhibited by 2 mM malate concentrations as applied in many SUIT protocols. This inhibition is less pronounced at higher succinate concentrations (10 mM up to 50 mM S). | :::: Recent studies showed that S- and NS-linked OXPHOS capacity is inhibited by 2 mM malate concentrations as applied in many SUIT protocols. This inhibition is less pronounced at higher succinate concentrations (10 mM up to 50 mM S). | ||
{{MitoPedia concepts | |||
|mitopedia concept=SUIT state | |||
}} | |||
Latest revision as of 18:29, 17 March 2022
Description
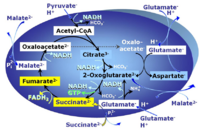
PGMS: Pyruvate & Glutamate & Malate & Succinate.
MitoPathway control state: NS-pathway control state
2-oxoglutarate is produced through the citric acid cycle from citrate by isocitrate dehydrogenase, from oxaloacetate and glutamate by the transaminase, and from glutamate by the glutamate dehydrogenase. If the 2-oxoglutarate carrier does not outcompete these sources of 2-oxoglutarate, then the TCA cycle operates in full circle with external pyruvate&malate&glutamate&succinate
Abbreviation: PGMS
Reference: Gnaiger 2020 BEC MitoPathways
PGMSL
- PGMS pathway in the LEAK state can be evaluated in the following SUIT protocols:
PGMSP
- PGMS pathway in the OXPHOS state can be evaluated in the following SUIT protocols:
- DL-Protocol for permeabilized fibers (pfi): SUIT-008 O2 pfi D014
- DL-Protocol for permeabilized cells (pce): SUIT-008 O2 ce-pce D025
PGMSE
- PGMS pathway in the ET state can be evaluated in the following SUIT protocols:
- DL-Protocol for isolated mitochondria and tissue homogenate (mt): SUIT-001 O2 mt D001
- DL-Protocol for permeabilized fibers (pfi): SUIT-001 O2 pfi D002
- DL-Protocol for permeabilized cells (pce): SUIT-001 O2 ce-pce D003
- DL-Protocol for permeabilized PBMC and PLT: SUIT-001 O2 ce-pce D004
- DL-Protocol for permeabilized fibers (pfi): SUIT-008 O2 pfi D014
- DL-Protocol for permeabilized cells (pce): SUIT-008 O2 ce-pce D025
Discussion
- Recent studies showed that S- and NS-linked OXPHOS capacity is inhibited by 2 mM malate concentrations as applied in many SUIT protocols. This inhibition is less pronounced at higher succinate concentrations (10 mM up to 50 mM S).
MitoPedia concepts:
SUIT state