Lima 2021 Nat Metab
| Lima A, Lubatti G, Burgstaller J, Hu D, Green AP, Di Gregorio A, Zawadzki T, Pernaute B, Mahammadov E, Perez-Montero S, Dore M, Sanchez JM, Bowling S, Sancho M, Kolbe T, Karimi MM, Carling D, Jones N, Srinivas S, Scialdone A, Rodriguez TA (2021) Cell competition acts as a purifying selection to eliminate cells with mitochondrial defects during early mouse development. Nat Metab 3:1091-108. https://doi.org/10.1038/s42255-021-00422-7 |
Lima A, Lubatti G, Burgstaller J, Hu D, Green AP, Di Gregorio A, Zawadzki T, Pernaute B, Mahammadov E, Perez-Montero S, Dore M, Sanchez JM, Bowling S, Sancho M, Kolbe T, Karimi MM, Carling D, Jones N, Srinivas S, Scialdone A, Rodriguez TA (2021) Nat Metab
Abstract: Cell competition is emerging as a quality-control mechanism that eliminates unfit cells in a wide range of settings from development to the adult. However, the nature of the cells normally eliminated by cell competition and what triggers their elimination remains poorly understood. In mice, 35% of epiblast cells are eliminated before gastrulation. Here we show that cells with mitochondrial defects are eliminated by cell competition during early mouse development. Using single-cell transcriptional profiling of eliminated mouse epiblast cells, we identify hallmarks of cell competition and mitochondrial defects. We demonstrate that mitochondrial defects are common to a range of different loser cell types and that manipulating mitochondrial function triggers cell competition. Moreover, we show that in the mouse embryo, cell competition eliminates cells with sequence changes in mt-Rnr1 and mt-Rnr2, and that even non-pathological changes in mitochondrial DNA sequences can induce cell competition. Our results suggest that cell competition is a purifying selection that optimizes mitochondrial performance before gastrulation.
• Bioblast editor: Gnaiger E
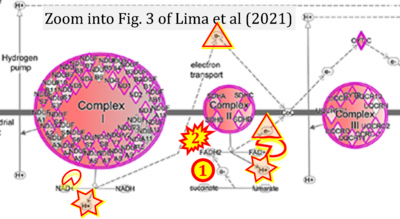
Correction: FADH2 and Complex II
- FADH2 is shown as the substrate feeding electrons into Complex II (CII). This is wrong and requires correction - for details see Gnaiger (2024).
- Gnaiger E (2024) Complex II ambiguities ― FADH2 in the electron transfer system. J Biol Chem 300:105470. https://doi.org/10.1016/j.jbc.2023.105470 - »Bioblast link«
Hydrogen ion ambiguities in the electron transfer system
Communicated by Gnaiger E (2023-10-08) last update 2023-11-10
- Electron (e-) transfer linked to hydrogen ion (hydron; H+) transfer is a fundamental concept in the field of bioenergetics, critical for understanding redox-coupled energy transformations.
- However, the current literature contains inconsistencies regarding H+ formation on the negative side of bioenergetic membranes, such as the matrix side of the mitochondrial inner membrane, when NADH is oxidized during oxidative phosphorylation (OXPHOS). Ambiguities arise when examining the oxidation of NADH by respiratory Complex I or succinate by Complex II.
- Oxidation of NADH or succinate involves a two-electron transfer of 2{H++e-} to FMN or FAD, respectively. Figures indicating a single electron e- transferred from NADH or succinate lack accuracy.
- The oxidized NAD+ is distinguished from NAD indicating nicotinamide adenine dinucleotide independent of oxidation state.
- NADH + H+ → NAD+ +2{H++e-} is the oxidation half-reaction in this H+-linked electron transfer represented as 2{H++e-} (Gnaiger 2023). Putative H+ formation shown as NADH → NAD+ + H+ conflicts with chemiosmotic coupling stoichiometries between H+ translocation across the coupling membrane and electron transfer to oxygen. Ensuring clarity in this complex field is imperative to tackle the apparent ambiguity crisis and prevent confusion, particularly in light of the increasing number of interdisciplinary publications on bioenergetics concerning diagnostic and clinical applications of OXPHOS analysis.
Labels:
Enzyme: Complex II;succinate dehydrogenase